By Ian Gormely
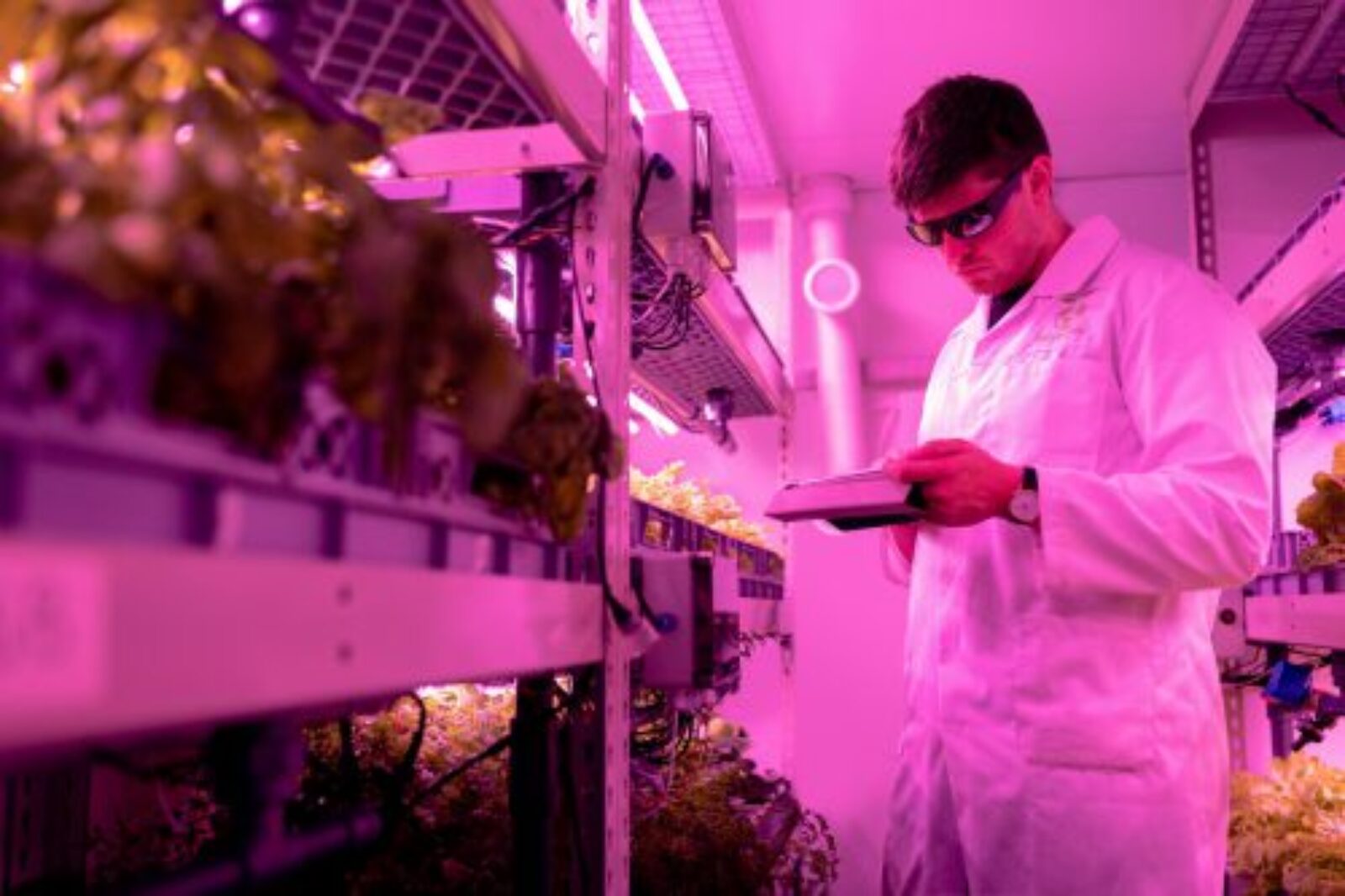
Data is the fuel that makes AI algorithms work. But good, usable data is not always easy to come by. It’s a problem that Vector Faculty Member Bo Wang knows all too well.
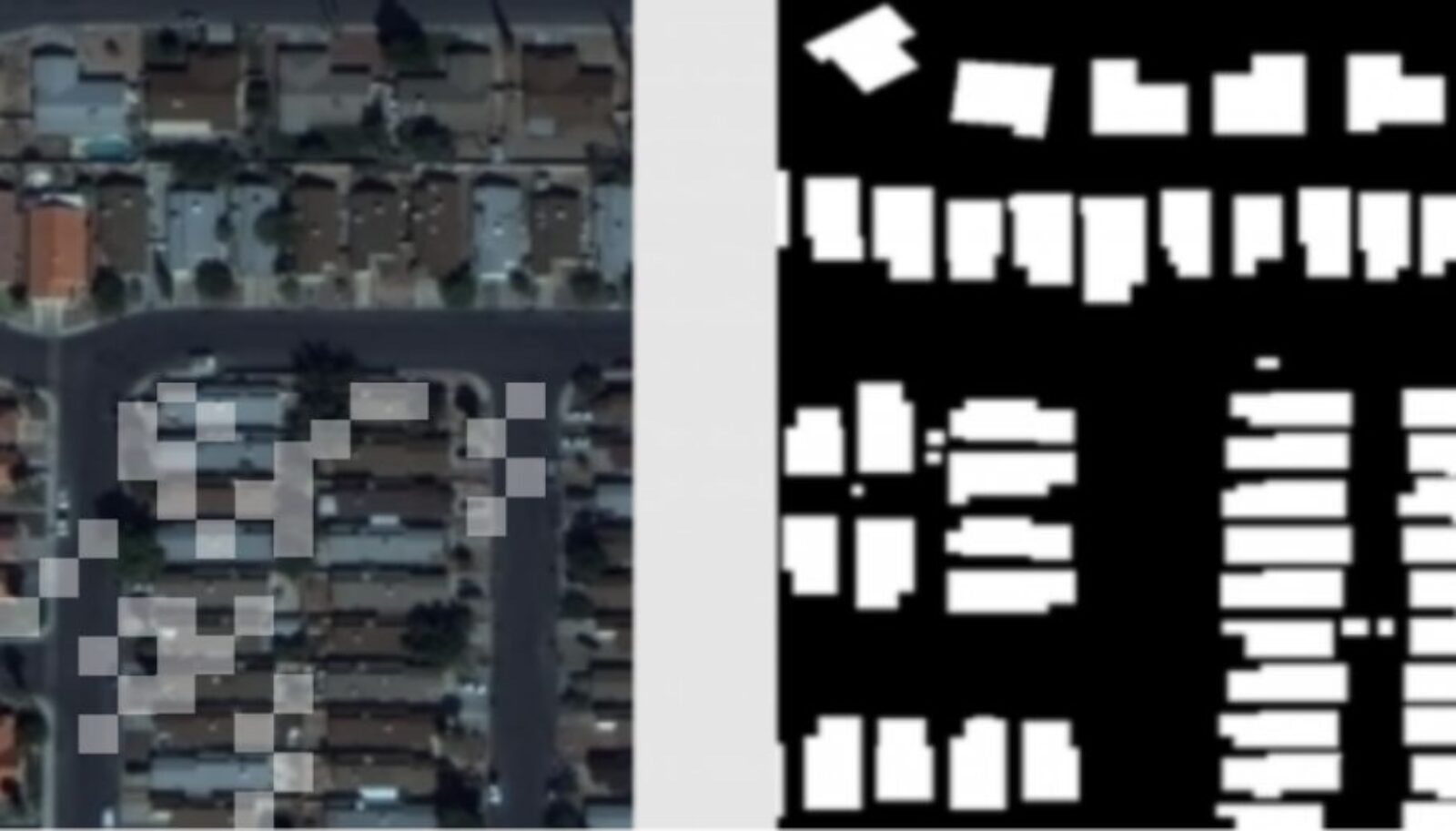
In order to build a set of medical data (such as medical images or clinical notes) big enough to make a machine learning model work, researchers like Wang often have to combine data from multiple hospitals. Specifically, Wang wanted to apply the classification model for cancer images collected from a group of hospitals to cardiac images. But the data hospitals collect, and the manner in which they collect and sort it, tends to vary from hospital to hospital and even department to department. Harmonizing them would be a difficult and time-consuming task. There had to be a better way.
Methods for targeting a single source domain adaptation – transferring one hospital’s model to another – already existed. But this “neglects more practical scenarios where training data are coming from multiple sources,” he says. He wondered: “Can we force the model to be generalizable to different domains?”
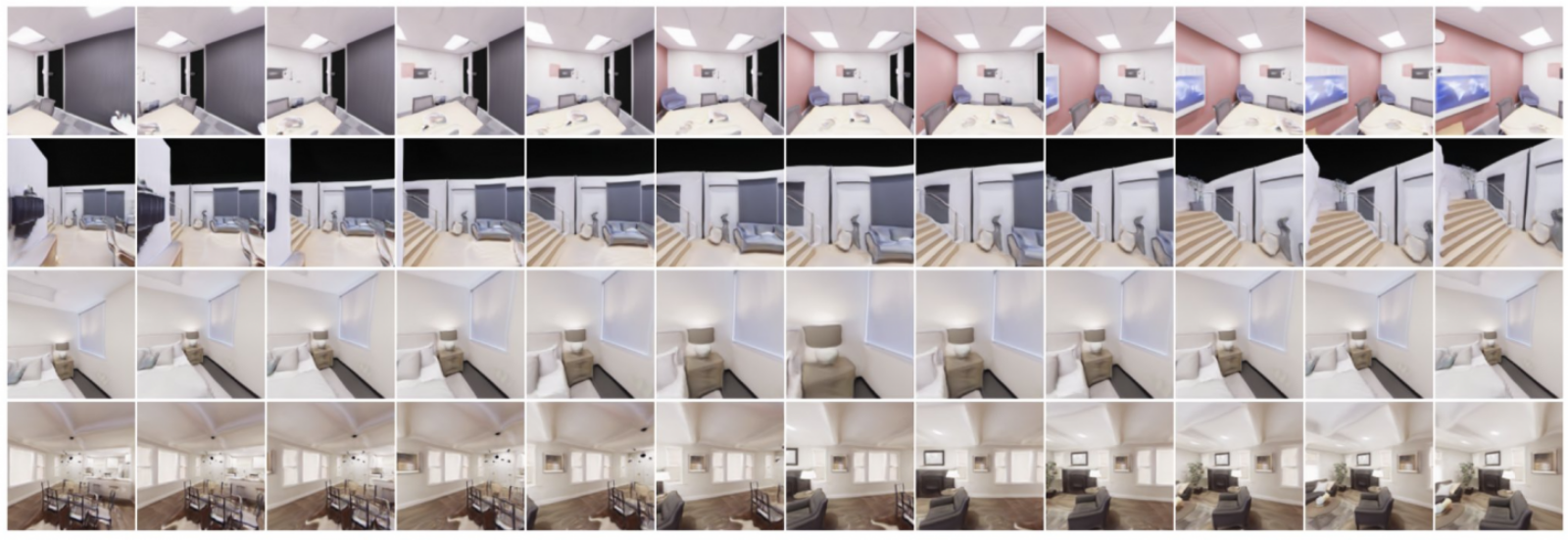
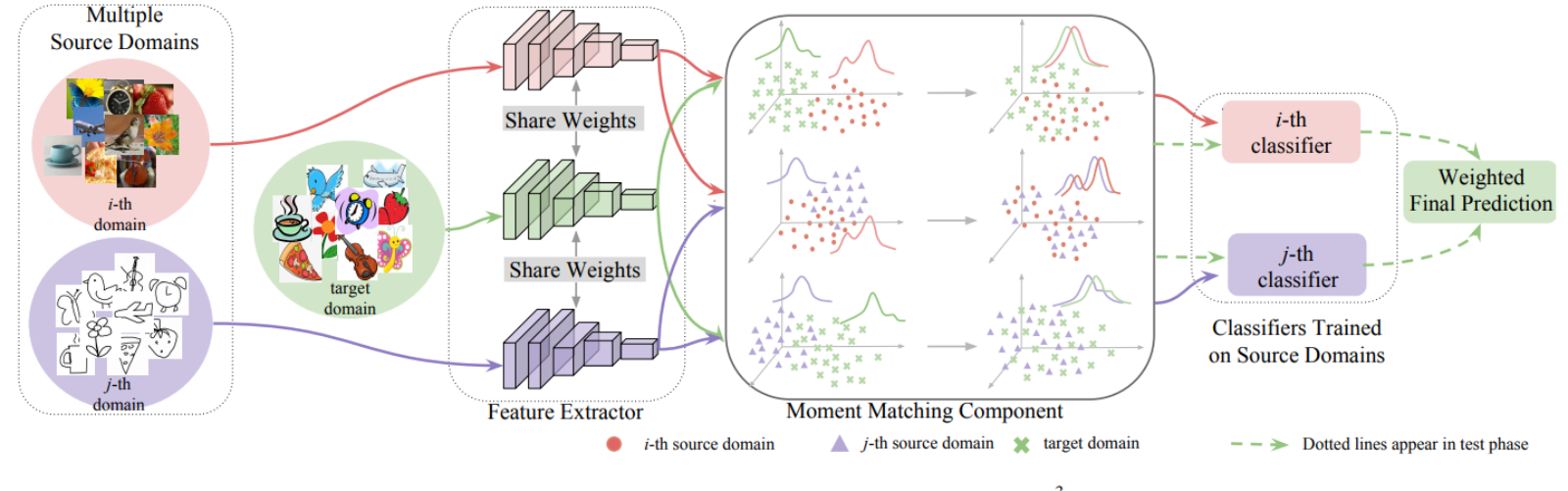
“Moment Matching for Multi-source Domain Adaption,” co-authored by Xingchao Peng, Qinxun Bai, Xide Xia, Zijun Huang, and Kate Saenko, and published in Proceedings of the IEEE International Conference on Computer Vision in 2019, suggests that yes, they very much can. In fact, their model – which they shorthand as M3SDA – not only offers a way to bring information from multiple sources into a single one, it also annotates unlabelled internet images, resulting in a data set of 600,000 images distributed across 345 different categories. “I believe it’s the largest collection of multiset domain adaptation,” says Wang.
And it’s already making an impact. Lauren Erdman, a Vector and SickKids Hospital PhD student under the supervision of Vector Faculty Member Anna Goldenberg, came across the paper while doing a literature review. “I immediately knew it would be useful in my work,” she says calling Wang’s approach “distinct” from others. Erdman is currently using’ Wang’s algorithm in two studies. The first uses M3SDA to harmonize differences in the data created by ultrasound machines. The other tweaks the algorithm — the outcome is a continuous-valued curve and alignment will be proportionate to curve similarity — to map the sound of uroflow (urination) to study its speed, volume and duration.
“We developed a very simple approach that can do a better job,” Wang says noting that while he and his team tested the model in computer vision, they also see “huge potential” for medical applications, particularly medical image analysis as well. “Most of the current medical image projects suffer from a lack of annotated images and the fact that models trained on data from one hospital cannot be directly applicable for images in the other hospital. Our method has the potential to conquer these two limitations at the same time.”