Vector research initiatives are making a remarkable impact in the field of public health and disease control.
By Natasha Ali
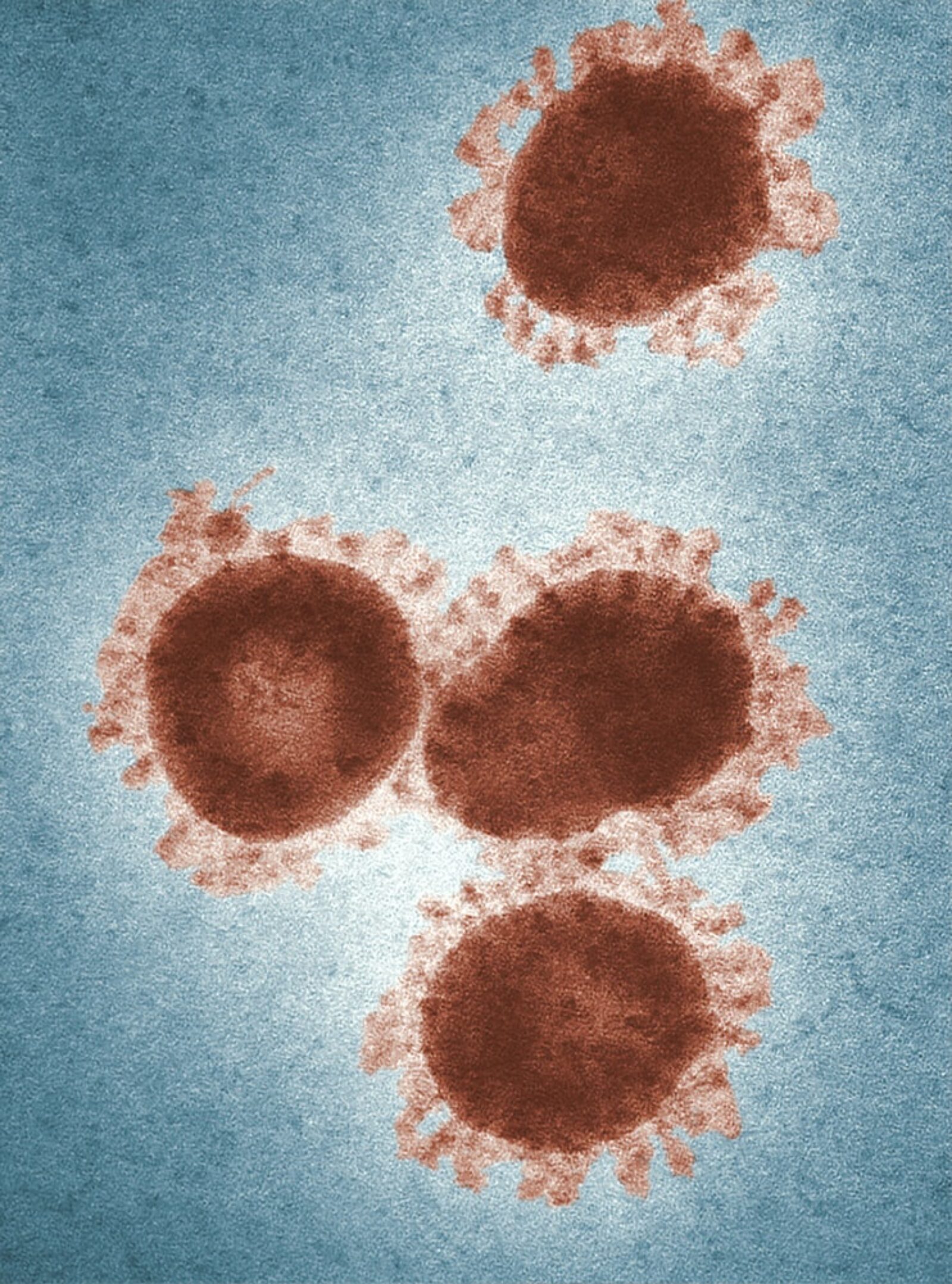
A new paper proposes a novel NLP model for infectious disease detection and monitoring. Co-authored by Vector Applied Machine Learning Scientist Shaina Raza, “Constructing a disease database and using natural language processing to capture and standardize free text clinical information,” uses COVID-19 knowledge as its basis and proposes an NLP approach to expedite the process of obtaining disease information from online sources and produce a structured disease database for clinical purposes.
Why are disease databases hard to build?
Through her extensive experience working with epidemiologists, public health experts, and medical researchers during the COVID-19 pandemic, Raza noticed a critical delay between the pace of clinical research and data analysis and the rate of viral mutation and disease spread.
“Everything was going so fast [especially with new variants of COVID],” says Raza, who works at the interface of AI and population health. “The research was, according to me, a little bit lagging behind.”
The ability to obtain health information in a timely manner is crucial for controlling large-scale outbreaks. Clinicians, epidemiologists, and public health experts rely on curated datasets from social media, electronic health records, and previous research findings to guide disease intervention and treatment strategies.
This meticulous data analysis process is usually carried out by specialized healthcare and infectious disease departments. By examining online and research-based sources, analysts can identify relevant disease information and put together clinical databases targeted for healthcare experts.
But challenges arise when trying to gather statistics about widespread diseases, like COVID-19, that are constantly developing and inflicting variable effects on individual patients. As a key aspect of disease intervention, clinical database management needs to be prioritized, but the lack of efficient structuring systems can stall the process.
To accelerate this intermediary step in time-sensitive situations, Raza’s research proposes an elaborate NLP model that can interpret free text from online sources such as blogs, social media posts, public forums, and medical notes and convert unstructured data into databases ready for clinical use.
What methodology was used for NLP model training and evaluation?
Conducting a thorough examination of medical case reports, published literature, and blogs, Raza, whose work generally focuses on applying AI and machine learning strategies to evaluate disease onset and expand public health initiatives, compiled a preliminary dataset of unstructured information. An annotation step was needed to label the dataset entities, forming the basis for machine training.
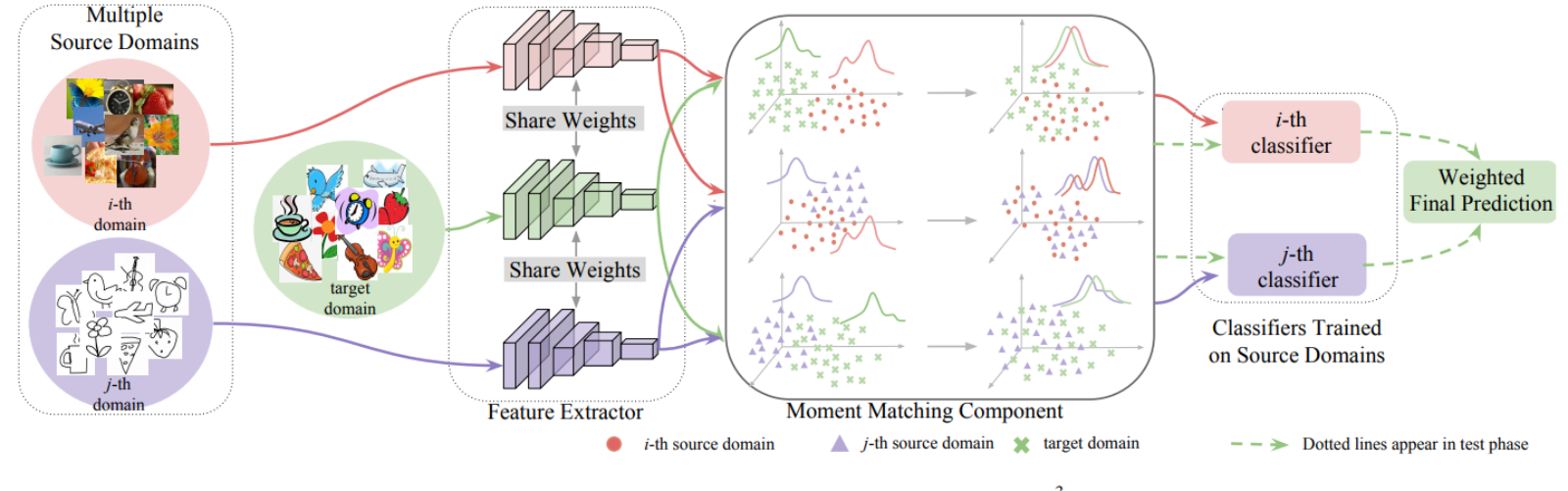
The extensive NLP framework included a named entity recognition model — a multilayer transformer-based model that trains deep neural networks on annotated sets of data, including symptoms, drug information, social determinants of health, risk factors, and disease dependencies. Using a subsequent relation extraction model, the NLP algorithm was able to infer connections between the corresponding entities and tabulate them based on clinical and non-clinical identifiers.
A two-phase evaluation method followed, wherein the proposed model was assessed against existing methods for disease detection and surveillance and a human evaluator was deployed to confirm the validity of the NLP framework as a whole.
What makes this research one of a kind?
The novelty of the study lies in the use of patient reports and case studies to evaluate non-clinical factors such as age, gender, race, geographical location, economic status, and other social determinants of health that could influence disease onset and symptom recognition.
Through implementing an adaptive NLP strategy to produce a structured dataset, medical professionals can extract key details about risk factors and treatment options as the disease evolves. The incorporation of non-clinical evaluation elements such as social determinants of health further solidifies the accuracy of the framework in identifying patterns and predicting medical outcomes.
That’s not to say that the NLP model will replace data analysts or eliminate jobs, nor will it supersede ongoing efforts, in the public health sector — this NLP methodology is only meant to improve pandemic surveillance and mitigate disease spread in public health emergencies.
“My motive is not to replace the data departments,” she says. “My motive is to take advantage of AI [to automate the data curation step] because it is something very much related to the lives of people.”
A caveat with this technology, she says, is the potential for machine error and inaccurate data linking, which dictates a constant need for human verification. “A human quality control loop should always be there, because sometimes, there is something that isn’t detected well by the AI model.”
What is the future of AI in clinical data analysis?
Discussing the potential uses of the NLP model for other diseases, Raza believes that by conducting periodic updates to the machine learning algorithms, the framework could be adapted to study other conditions and future outbreaks.
“In terms of generalizability, this particular dataset was prepared in relation to COVID-19, and long COVID. However, this framework can be used for other diseases, but data has to be rebuilt,” she says. “If somebody wants to do research on cancer, diabetes, or hepatitis, then they have to find the data that belongs to these particular diseases.”
Still in its experimental phase, Raza hopes that this novel NLP framework could be deployed for use in hospitals and public health organizations to consolidate database analysis and disease monitoring. But, she says, the largest hurdle is garnering enough computational resources, multidisciplinary support and necessary backing to train the NLP algorithms, and recruiting machine learning experts in healthcare settings.
“It is not a simple task,” she adds. “If for example, some organization wants to deploy it, they will need some retraining environment and some deployment environment. But there definitely is a plan.”