Vector researchers developed CRISPNAM-FG, a trustworthy AI model that predicts the risk of developing diabetes-related foot complications for patients discharged from hospitals while providing complete transparency in how each decision is made – addressing the critical need for interpretable AI in health care where understanding model reasoning is paramount for clinical adoption. This breakthrough demonstrates that AI systems can achieve high accuracy while maintaining full interpretability, solving a fundamental challenge that has limited the deployment of AI in high-stakes medical settings. Access the full paper here.
TLDR: Uncover groundbreaking AI research in 3 minutes
This concise summary bridges the gap between complex scientific advancements and everyday understanding. Ideal for enthusiasts and non-researchers, start listening now.
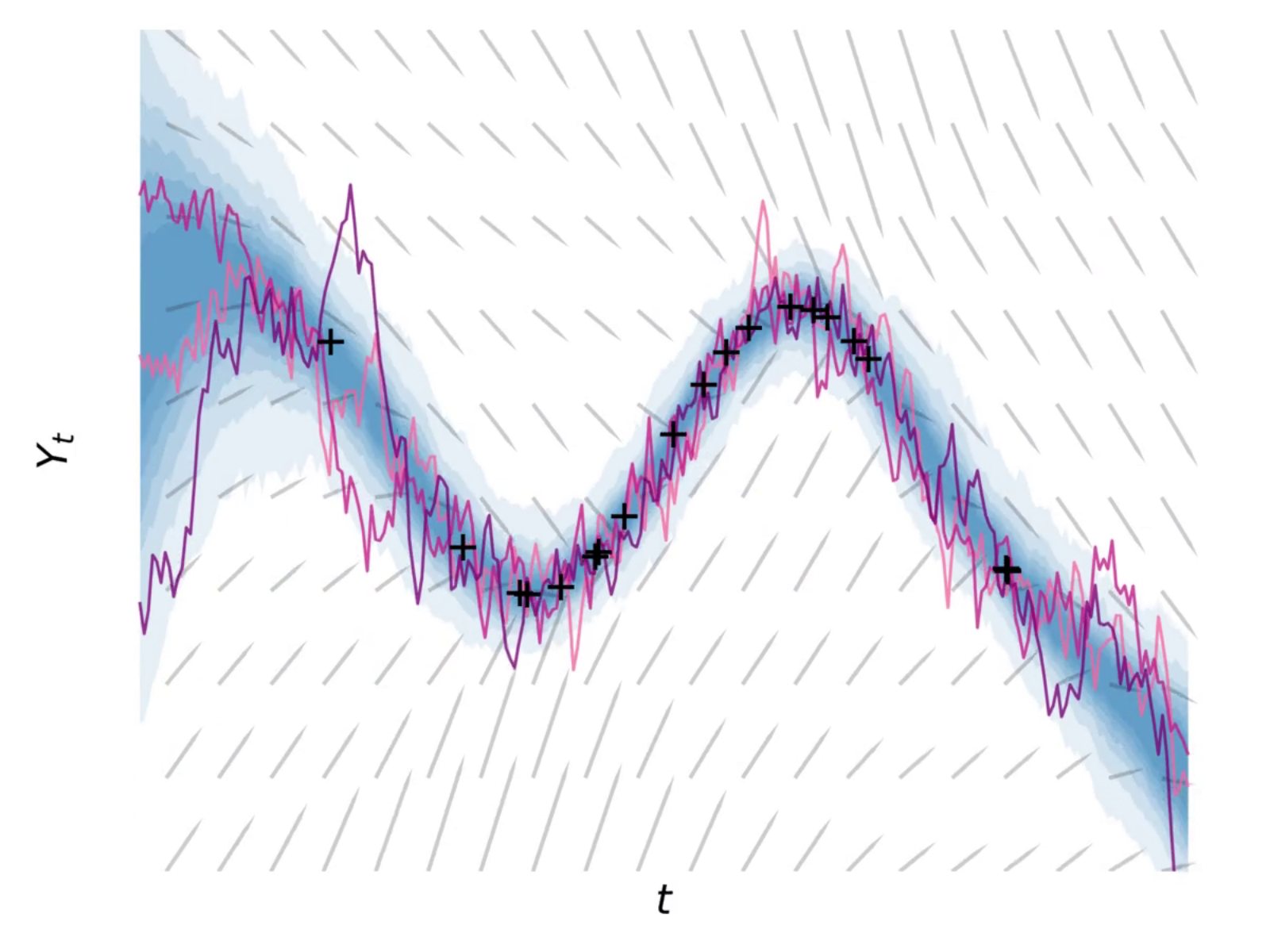
Survival analysis is a statistical approach for analyzing time-to-event data and it involves essentially, studying how long it takes until something happens (like disease onset, recovery, or death) and what factors influence that timing.Traditional survival analysis focuses on predicting risk for a single event, but many real-world health care scenarios involve multiple “competing” events where experiencing one prevents others from occurring. For people with diabetes, this might include the risk of foot complications competing with risk of mortality from other causes.
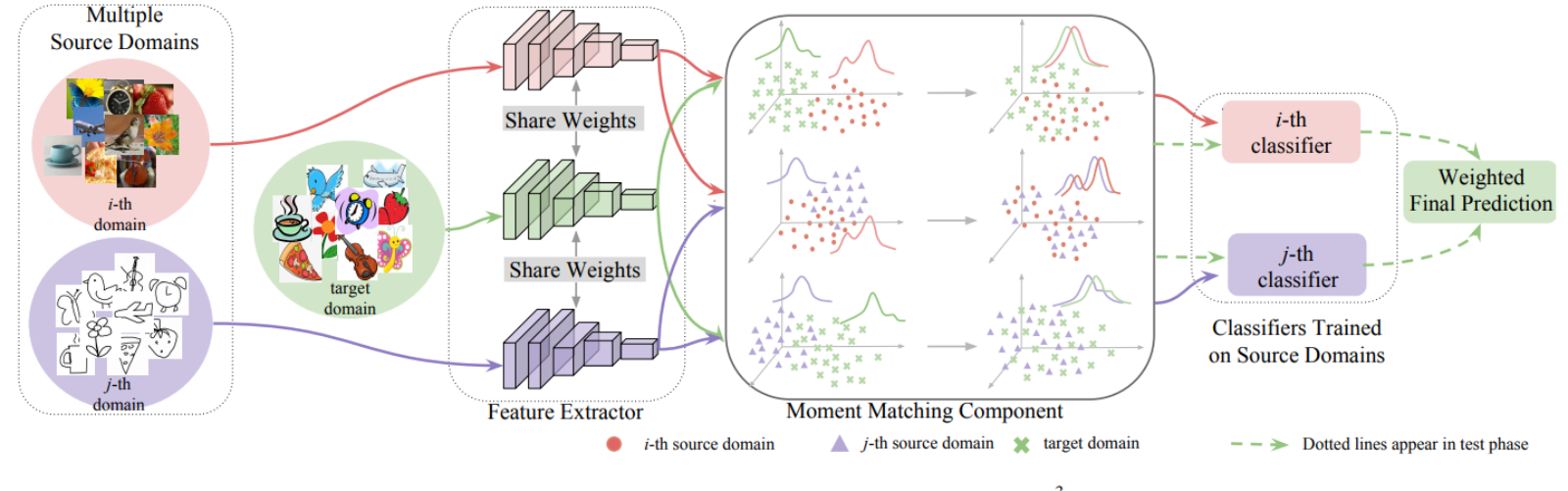
Recent deep learning advances like DeepHit and DeepSurv have improved predictive performance for competing risks scenarios, but these gains come at the cost of interpretability, creating black-box models that are difficult for clinicians to trust and validate in practice.
Our team’s research goal is to enable accurate and transparent prediction of time-to-event outcomes where multiple risks coexist, while maintaining a level of interpretability that supports clinical trust, validation, and policy decision-making. This work demonstrates that high predictive performance and intrinsic interpretability can coexist in competing risks survival analysis for real-world health care application.
This work represents a collaborative effort between Vector Institute, GEMINI, Unity Health, and Diabetes Action Canada, bringing together methodological expertise, large-scale hospital data, and domain knowledge in diabetes care. Together, the research team developed CRISPNAM-FG (Competing Risks Survival Prediction using Neural Additive Models: Fine Gray), an intrinsically interpretable survival model that handles competing risks using the Fine-Gray formulation while maintaining both the predictive power of deep learning and the interpretability of classical statistical models.
The interpretability challenge in survival analysis
Model interpretability is crucial for establishing AI safety and clinician trust in medical applications. Recent deep learning models have achieved very good predictive performance, but their limited transparency as black-box models hinders their integration into clinical practice.
Existing deep survival approaches for competing risks, including DeepHit and Neural Fine-Gray, lack inherent interpretability, particularly at the feature level. This opacity makes it challenging to understand individual feature contributions to risk predictions across different competing events.
Post-hoc explanation methods like LIME and SHAP have limitations. They generally suffer from low fidelity, leading to unfaithful or misleading explanations and can be easily fooled. Unlike black-box deep survival models, the CRISPNAM-FG approach is intrinsically interpretable and does not rely on post-hoc explanation methods, which are often unstable and difficult to validate in health care settings.
CRISPNAM-FG architecture: Building interpretability from the ground up
The key contribution is the integration of the Fine-Gray competing risks framework with Neural Additive Models in a single, end-to-end trainable architecture. Each clinical variable is modeled independently using a small neural network, allowing the system to capture non-linear effects while preserving feature-level transparency.
Neural Additive Model foundation
Following the Neural Additive Model framework, each input feature is processed by its own dedicated neural network, called a FeatureNet. These feature-specific sub-networks learn the non-linear contribution of each individual feature to the overall risk score while preserving interpretability by isolating feature effects.
Each FeatureNet takes a scalar input and produces a hidden representation through fully-connected feedforward neural networks with hyperbolic tangent activation functions.
Risk-specific projections
The model allows each patient characteristic to influence different outcomes in different ways. For example, high blood sugar might strongly increase the risk of foot complications while having a different effect on mortality risk. This flexibility is crucial because the biological mechanisms driving each outcome can be distinct.
For each variable and each competing outcome, the model learns separate transformations that capture how that variable contributes to risk. To ensure fair comparisons across different outcomes, these transformations are standardized to the same scale. This prevents one outcome from appearing artificially more important simply due to differences in measurement units or mathematical scaling.
Fine-Gray formulation
The Fine-Gray approach models the probability of experiencing a specific event over time, accounting for the presence of competing events. Unlike traditional methods that remove patients from consideration once they experience any event, Fine-Gray keeps tracking patients even after they’ve had a competing event. This provides a more realistic picture of risk in clinical settings where multiple outcomes can occur.
The model calculates risk by combining a baseline hazard that changes over time with patient-specific factors. It then converts this into cumulative incidence: the probability that a patient will experience the event of interest by a given time point.
Interpretability through shape functions
The model’s key strength is its built-in interpretability. Rather than producing predictions without explanation, it generates “shape functions” that visualize how each patient characteristic influences risk.
For each clinical variable (like age, blood glucose, or blood pressure), the model creates a curve showing how different values of that variable affect risk, separately for each possible outcome. For example, you might see how increasing HbA1c levels progressively increase the risk of foot complications versus mortality.
The system provides two key interpretability features:
- Shape function plots that show how each variable influences risk across its entire range of values.
- Feature importance rankings that identify which factors most strongly drive each competing outcome
This transparency allows clinicians to understand not just who is at high risk, but why the model reaches that conclusion, making the predictions actionable and verifiable against clinical knowledge.
Real-world validation: Diabetic foot complications in Ontario
The clinical problem and GEMINI dataset
The researchers applied CRISPNAM-FG to predict future diabetic foot complications after hospital discharge, using data from over 100,000 patients across 29 hospitals in Ontario. Foot complications are treated as the primary outcome, with in-hospital death modeled as a competing risk.
The dataset, obtained from GEMINI, included all adult patients with diabetes discharged home between April 2016 and March 2023. A foot complication was defined as the first hospitalization for foot ulcer, infection, gangrene, or amputation. After applying exclusion criteria, the final cohort comprised 107,386 patients.Patient characteristics included a median age of 72.0 years, with 46.2% female patients. The cohort had significant comorbidity burden, with 39.8% having a Charlson Comorbidity Index1 of 1, and 28.4% having an index of 2 or higher.
Model performance
CRISPNAM-FG demonstrated strong discriminative performance on the GEMINI dataset, achieving competitive results compared to existing deep competing risks models. We evaluated the model using two standard measures. TD-AUC (time-dependent area under the curve) measures how well the model distinguishes between patients who will and won’t experience an event by a given time point, with values closer to 1.0 indicating better discrimination. TD-CI (time-dependent concordance index) similarly assesses whether the model correctly ranks patients by their risk, with higher values indicating more accurate risk ordering.
For foot complications (Risk 1), the model achieved TD-AUC values of 0.766 ± 0.042 at the 25th percentile, 0.763 ± 0.047 at the median, and 0.763 ± 0.052 at the 75th percentile. For the competing mortality risk (Risk 2), TD-AUC values were 0.739 ± 0.057, 0.726 ± 0.058, and 0.707 ± 0.060 across the same percentiles.
The model demonstrated strong discriminative performance compared to existing deep competing risks models, while producing clinically coherent risk patterns that align with medical knowledge.
Clinical insights from interpretable shape functions
Shape functions are the core interpretability mechanism in CRISPNAM-FG. These are learned curves that show how each individual feature influences the model’s prediction across its range of values. Unlike traditional linear models where each feature has a single coefficient, shape functions can capture non-linear relationships. For example, they might show that risk increases slowly at low HbA1c levels but accelerates rapidly at higher values. Each feature gets its own shape function, and the final prediction is simply the sum of all these individual feature contributions. This additive structure is what makes CRISPNAM-FG interpretable: clinicians can examine each shape function independently to understand exactly how the model uses each variable to calculate risk.
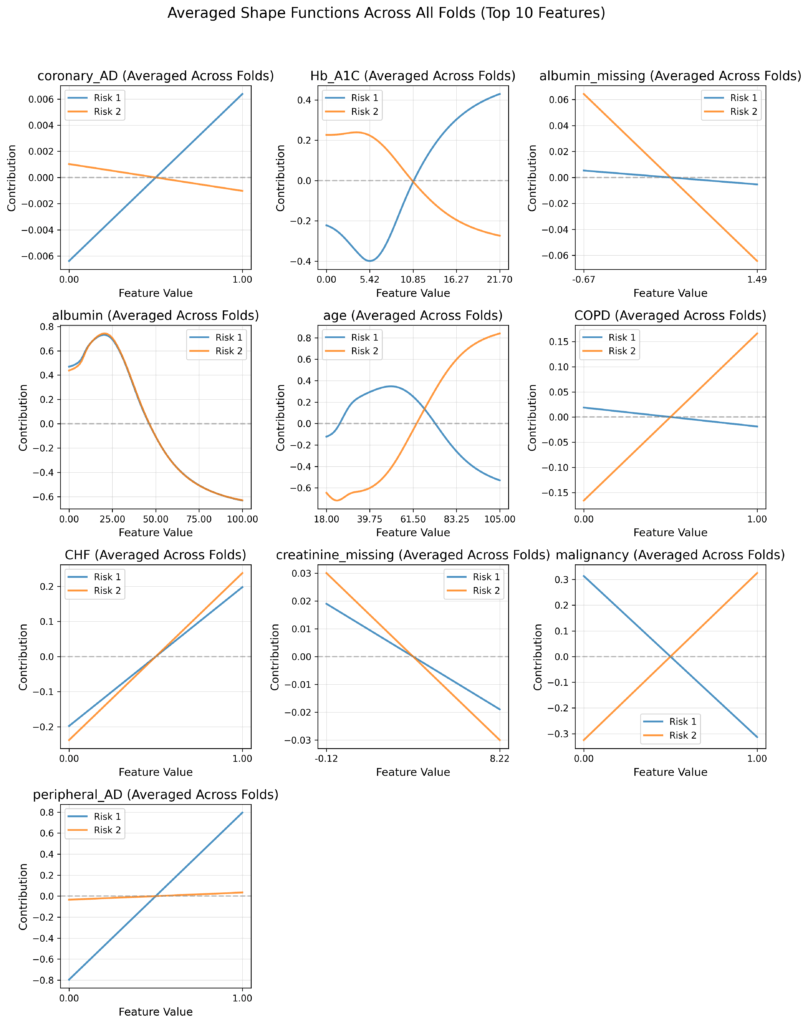
The shape functions computed from the trained CRISPNAM-FG model on the GEMINI dataset revealed clinically meaningful and sometimes counterintuitive patterns that demonstrate the model’s ability to capture complex clinical relationships:
- Glycemic control (HbA1c): The shape functions show a fascinating pattern where higher HbA1c values increase foot complication risk as expected, but actually decrease mortality risk. This counterintuitive finding demonstrates the model’s ability to reveal complex clinical relationships, specifically that of patients with poor glycemic control may be more likely to develop foot complications before experiencing fatal events.
- Cardiovascular comorbidities: The presence of coronary artery disease shows increased contribution to foot complication risk, consistent with established medical knowledge about the relationship between cardiovascular disease and diabetic complications. Congestive heart failure (CHF) demonstrates an increasing contribution to both foot complications and mortality, showing how cardiovascular comorbidities affect multiple outcomes.
- Peripheral vascular disease: The shape functions show a strong, almost linear increase in foot complication risk with peripheral artery disease, while showing minimal impact on mortality risk. This aligns perfectly with clinical understanding that peripheral vascular disease directly affects limb perfusion and wound healing.
- Age patterns: Age exhibited distinct patterns for the two outcomes. The risk contribution for foot complications increases until approximately 60 years before declining, whereas mortality risk increases monotonically with age. This divergence after age 60 likely reflects competing risks, where older patients succumb to systemic complications before developing foot-specific problems.
- Nutritional status (Albumin): Lower albumin levels contribute to both risks, but the relationship is more pronounced for mortality risk, reflecting the known association between hypoalbuminemia and poor outcomes. The shape functions show albumin’s protective effect diminishing as levels drop below normal ranges.
- Disease-specific effects: The model correctly identified that chronic obstructive pulmonary disease (COPD) does not contribute substantially to foot complication risk but significantly increases mortality risk. Similarly, malignancy showed stronger association with mortality than with foot complications. These distinctions are essential for appropriate post-discharge planning.
- Missing data patterns: Missing values for creatinine and albumin indicated selective test ordering, where clinicians did not order these tests in the absence of clinical suspicion for renal dysfunction or hypoalbuminemia-related complications.
Supporting preventive care and resource allocation
CRISPNAM-FG is designed specifically to support preventive care and resource allocation. By identifying patients at high risk of foot complications, health systems can prioritize early screening, outpatient follow-up, and multidisciplinary interventions that are known to reduce amputations and downstream costs.
Because the model is interpretable, its outputs can be reviewed, audited, and communicated to clinicians, patients, and policymakers. The transparency enables health care providers to understand not just which patients are at risk, but why they are at risk and which specific factors drive their elevated risk profile.
This interpretability is crucial for clinical adoption because clinicians can validate the model’s reasoning against their clinical experience and use the detailed risk attribution to guide targeted interventions. For instance, a patient flagged primarily for poor glycemic control would benefit from different interventions than one flagged for vascular disease.
Model advantages and limitations
CRISPNAM-FG provides several key advantages over existing approaches:
- Intrinsic interpretability: Unlike black-box models, CRISPNAM-FG provides transparent reasoning through shape functions and feature importance plots that show exactly how each feature contributes to each competing risk.
- Risk-specific insights: The model enables the same variable to have different effects on different outcomes, revealing complex clinical relationships that would be obscured in single-outcome models.
- Clinical coherence: The shape functions produce clinically meaningful patterns that align with established medical knowledge while revealing new insights about risk interactions.
- Individual-level transparency: The model enables understanding of feature contributions for individual patients, not just global feature importance.
We do acknowledge that our model does not capture feature-level interactions beyond additive effects, which may limit its ability to model complex relationships. Additionally, while discrimination performance is competitive, the model’s calibration performance lags behind some alternatives like DeepHit.
Real-world implementation pathway
While CRISPNAM-FG has been evaluated using retrospective data and has not yet been deployed in routine clinical practice, this work builds on the team’s proven track record in clinical implementation. As part of the same broader initiative, the research team has previously deployed an earlier Fine-Gray subdistribution model at St Michael’s Hospital.
This implementation experience demonstrates the practical feasibility of interpretable survival models in real-world clinical settings and provides a pathway for scaling CRISPNAM-FG to broader health care applications.
Conclusions and future directions
The researchers demonstrate that it is possible to achieve competitive predictive performance in competing risks survival analysis without sacrificing interpretability. CRISPNAM-FG offers a transparent, clinically aligned approach to risk prediction that is well suited for real-world health care deployment and population-level decision-making.
The model produces competitive discrimination scores while providing transparency through shape functions that reveal detailed feature contributions to each risk. The application to diabetic foot complications shows the model’s practical utility in real-world health care scenarios, revealing clinically meaningful insights about risk factors and their differential impacts on competing outcomes.
Future research directions include investigating temporal feature networks that can capture time-varying feature contributions for each risk and developing improved loss functions to enhance the model’s calibration performance.
The core value proposition for health care applications is the inherent interpretability that enables clinicians to understand, validate, and trust model predictions which is a crucial requirement for safe and effective AI integration in clinical practice. Through this collaborative effort between Vector Institute, GEMINI, and Diabetes Action Canada, the research team has demonstrated that the future of health care AI lies not in choosing between accuracy and interpretability, but in architecting both into unified, clinically valuable systems.

Created by AI, edited by humans, about AI
This blog post is part of our ‘ANDERS – AI Noteworthy Developments Explained & Research Simplified’ series. Here we utilize AI Agents to create initial drafts from research papers, which are then carefully edited and refined by our humans. The goal is to bring you clear, concise explanations of cutting-edge research conducted by Vector researchers. Through ANDERS, we strive to bridge the gap between complex scientific advancements and everyday understanding, highlighting why these developments are important and how they impact our world.
Additional notes
1 Charlson Comorbidity Index is a scoring system that quantifies a patient’s overall disease burden by assigning points for the presence of specific chronic conditions (like diabetes, heart disease, or cancer), with higher scores indicating more severe comorbidity and greater health risk.