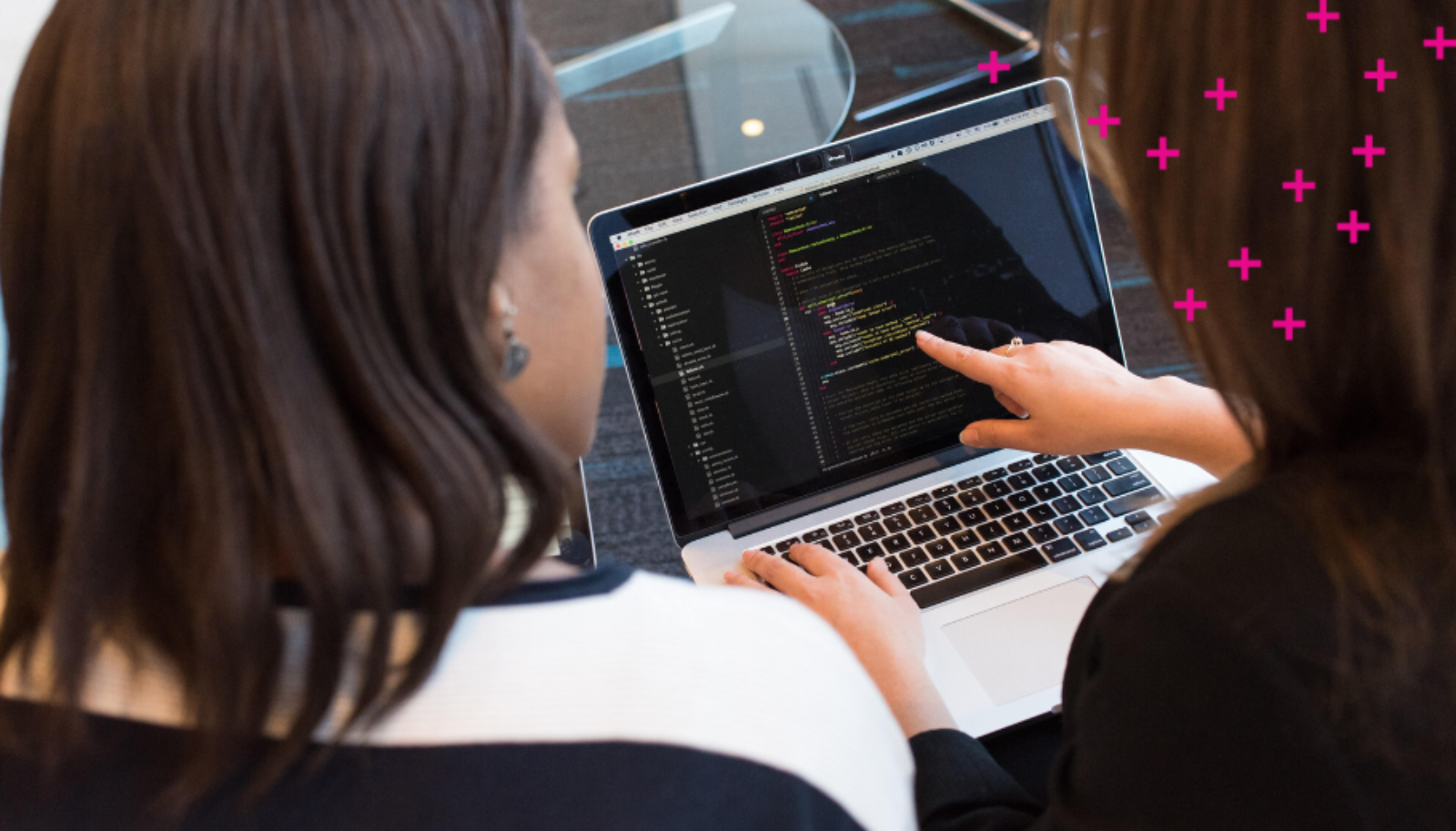
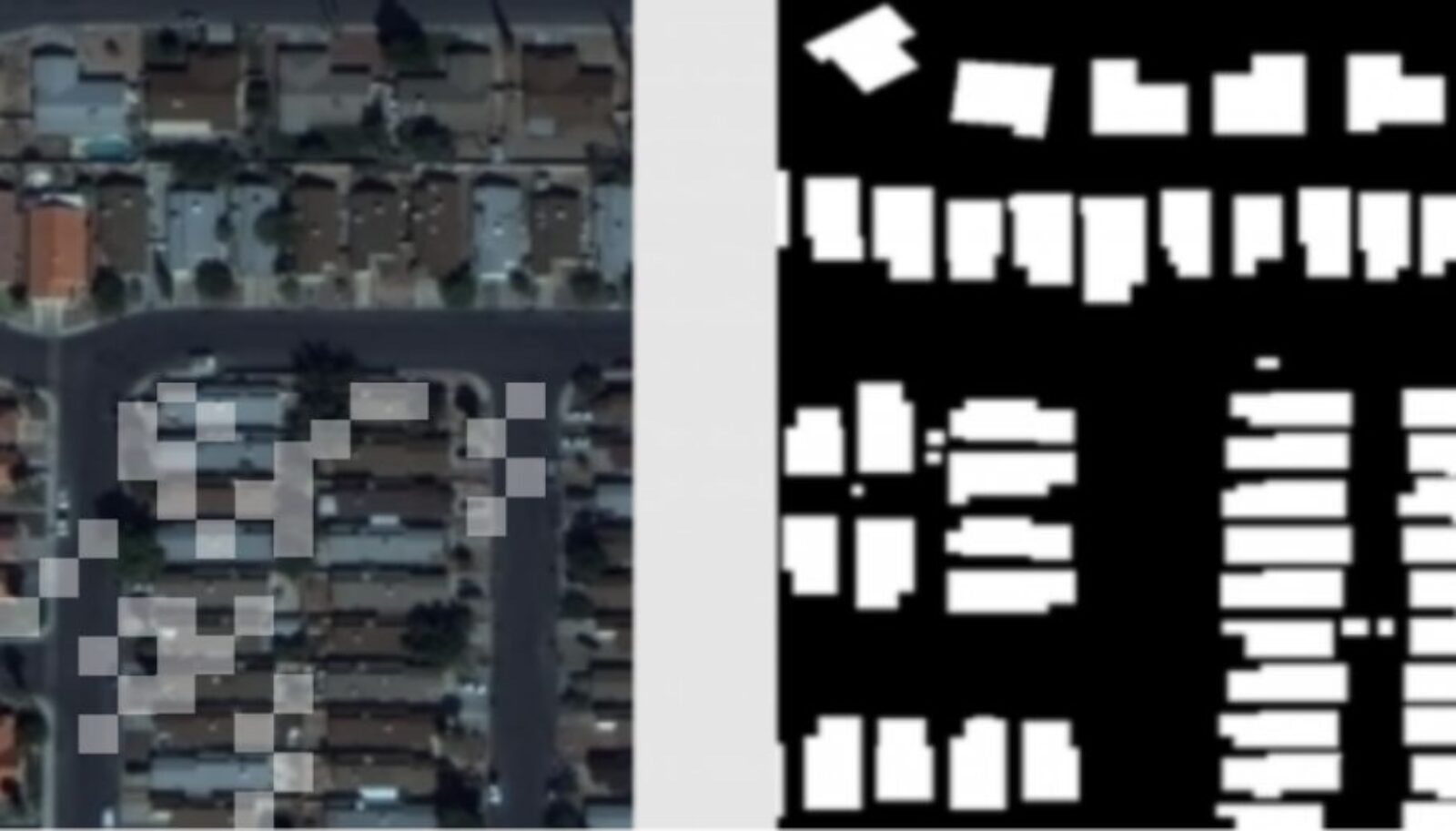
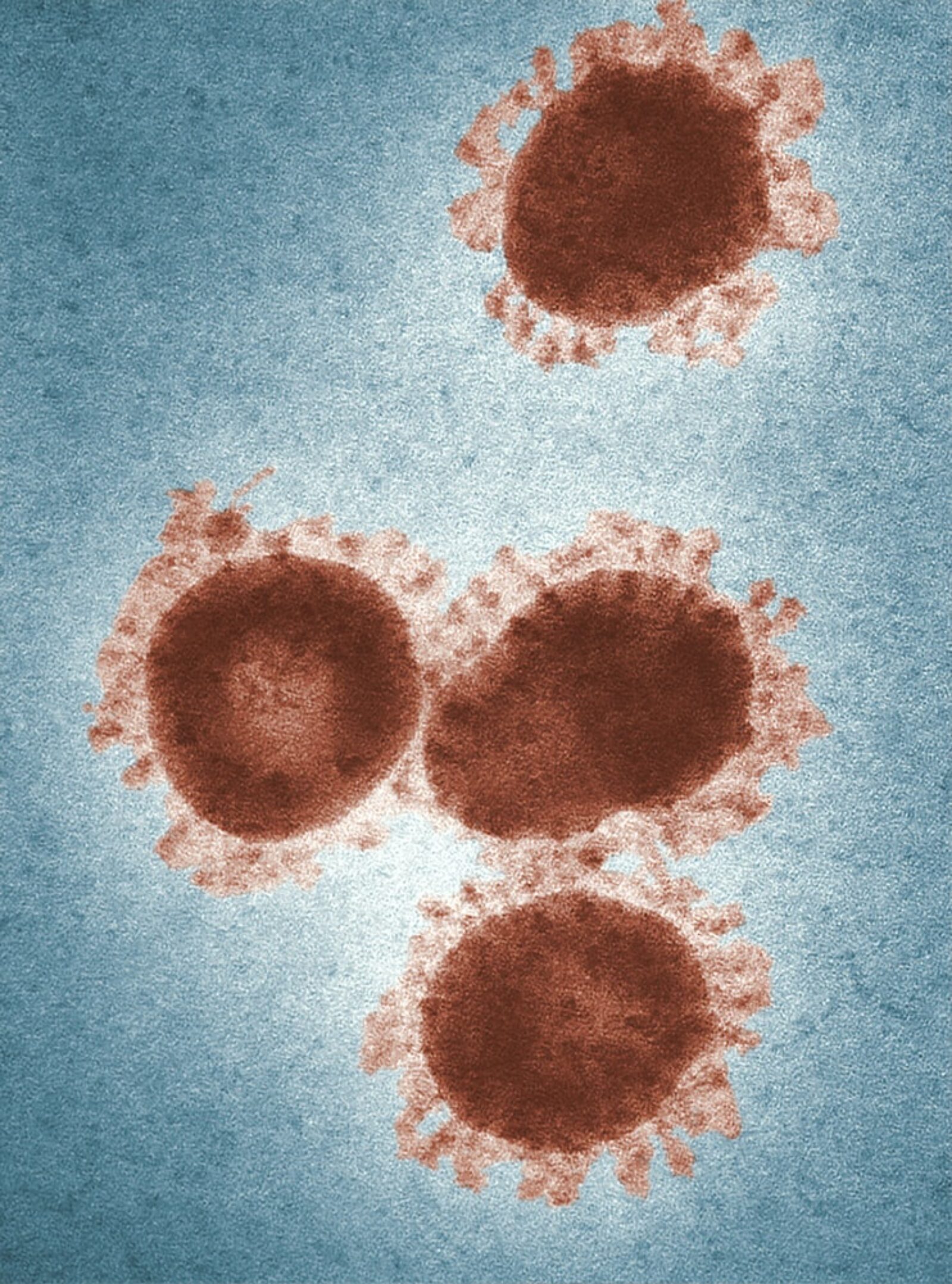
Researchers have developed an AI-enabled genotyping tool that can analyze viral genome data from a nasal swab. Notably, it can be used to determine different variants of concern (VOC) of COVID-19.
The tool was developed by Vector researcher Bo Wang and his team as part of the Viral AI network developed by Ontario-based tech company DNAStack and the COVID Cloud Supercluster Project. “Our tool takes large amounts of viral genomes data as a reference and uses ML to learn a useful representation of viral genomes,” says Wang.
Viral AI is powered by software developed with support from the federal COVID Cloud Supercluster Project. It offers an alternative to the centralized model, where data is uploaded to a single database. Instead, it uses a federated architecture to connect, analyze, and share data without moving it allowing for faster, more efficient, regulatory
compliant, and regionally sovereign data management.
The Province of Ontario is currently working with DNAStack to create a database that organizes genomic data and a platform that visualizes the genomic data in real-time for VOC genotyping. Once established these digital tools will help improve public health officials’ capacity to identify VOCs and aid in the government’s pandemic planning response.
Wang and his team are currently only working with COVID-19 data, but he says their tool could be repurposed for any other virus, like the common flu.